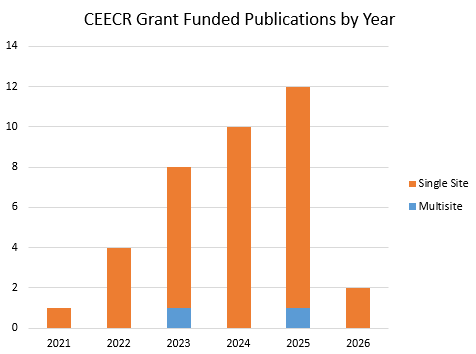
Graph includes preprints – last update 05/19/2026
Bisson, W. H., Currie, R. A., Lim, E. L., Mlynarczyk, C., Tokar, E. J., Colacci, A., & Colacino, J. A. (2026). Rethinking the microenvironment’s role in chemical-induced Malignancy. Toxicological sciences: an official journal of the Society of Toxicology, 209(2).
Abstract
Why and how does Cancer start? Building from a Symposium at the 2025 Society of Toxicology meeting, we convened a group of international experts to answer this seemingly simple question. As experimental evidence has evolved, perspectives on cancers’ origins have shifted from the accumulation of DNA mutations in single cells to complex processes involving signals from an altered tissue microenvironment which promote tumorigenesis. Carcinogen exposures impact the biology of the microenvironment in complex and tissue-specific ways. These changes can include the infiltration of inflammatory cells that produce growth factors, neo-angiogenesis, morphological changes, and immune tolerance that avoids immune-mediated elimination. In this in-depth review, we discuss the evidence linking chemical-driven microenvironmental changes in the development of a range of solid and liquid tumors. We discuss specific phenotypic alterations, such as selection pressure driving clonal expansion and cellular plasticity and reacquisition of stem cell states, linked to carcinogen-induced changes in the microenvironment. We describe assays and biomarkers which can allow us to experimentally assess links between chemical exposures, the microenvironment, and Cancer phenotypes. We end by discussing how understanding the role of the microenvironment and Malignancy in toxicology is essential for accurate Cancer hazard evaluation, development of next-generation risk assessment frameworks, identifying new strategies for Cancer prevention, and improving patient care.
Keywords: biomarkers; Cancer prevention; Cancer risk assessment; microenvironment; promotion.
Published by Oxford University Press on behalf of the Society of Toxicology 2025.
Kundu, R., Shi, X., Kleinsasser, M., Fritsche, L. G., Salvatore, M., & Mukherjee, B. (2026). A doubly robust framework for addressing outcome-dependent selection bias in multi-Cohort EHR studies. Biostatistics (Oxford, England), 27(1), kxag001. https://doi.org/10.1093/biostatistics/kxag001
Abstract
Selection bias can hinder accurate estimation of association parameters in binary disease risk models using non-probability samples like electronic health records (EHRs). The issue is compounded when participants are recruited from multiple clinics/centers with varying selection mechanisms that may depend on the disease/outcome of interest. Traditional inverse-probability-weighted (IPW) methods, based on constructed parametric selection models, often struggle with misspecifications when selection mechanisms vary across cohorts. This paper introduces a new Joint Augmented Inverse Probability Weighted (JAIPW) method, which integrates individual-level data from multiple cohorts collected under potentially outcome-dependent selection mechanisms, with data from an external probability sample. JAIPW offers double robustness by incorporating a flexible auxiliary score model to address potential misspecifications in the selection models. We outline the asymptotic properties of the JAIPW estimator, and our simulations reveal that JAIPW achieves up to 6 times lower relative bias and 5 times lower root mean square error (RMSE) compared to the best performing joint IPW methods under scenarios with misspecified selection models. Applying JAIPW to the Michigan Genomics Initiative (MGI), a multi-clinic EHR-linked biobank, combined with external national probability samples, resulted in Cancer-sex association estimates closely aligned with national benchmark estimates. We also analyzed the association between Cancer and polygenic risk scores (PRS) in MGI to illustrate a situation where the exposure variable is not measured in the external probability sample.
Keywords: Data integration; double robustness; inverse probability weighting (IPW); joint augmented inverse probability weighted (JAIPW); multi-Cohort sampling; selection bias.
© The Author(s) 2026. Published by Oxford University Press. All rights reserved. For commercial re-use, please contact [email protected] for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact [email protected].
Abstract
Background and aims: Per- and poly-fluoroalkyl substances (PFAS) are persistent chemicals that contaminate air, water, soil, and food. Due to their widespread use, PFAS are detectable in most of the US population, raising concerns about potential health impacts, including a possible association with colorectal Cancer (CRC). We conducted a scoping review of previously published studies to consolidate the current understanding of PFAS and its effect on CRC risk, identify knowledge gaps, and propose future directions for research. Methods: We systematically searched PubMed, Scopus, and Web of Science for studies published through December 2024 that examined PFAS exposure in relation to CRC risk or progression. Data were independently extracted using a standardized protocol and structured according to a predefined Population, Exposure, Comparator, Outcome (PECO) framework. To evaluate internal validity, studies were categorized by evidence stream (in vitro, animal, and epidemiological) and assessed using established quality appraisal criteria. Results: Twenty-six studies were identified, including 5 reviews, 3 in vitro studies, 6 animal studies, and 12 epidemiological studies. In vitro research consistently demonstrates that PFAS exposure promotes CRC cell proliferation and migration, highlighting key mechanistic pathways. However, findings from animal and epidemiological studies are mixed, with some indicating increased intestinal tumorigenesis while others report null or protective effects. Notably, major studies were cross-sectional, retrospective, or ecological, emphasizing the need for high-quality longitudinal research to clarify PFAS’s role in CRC risk and progression. Conclusion: Evidence on the relationship between PFAS exposure and CRC risk remains limited and inconclusive. Large-scale, prospective epidemiological studies that incorporate biomarker-based exposure assessment, including pre-diagnostic PFAS levels, diverse populations, and rigorous study design are needed to clarify the role of PFAS in CRC development. Such research could provide the role of PFAS in CRC development and progression, guiding public health policies and support targeted interventions to reduce CRC risk.
Keywords: Animal Studies; Colorectal Cancer; Epidemiological Studies; In Vitro Studies; Per- and Polyfluoroalkyl Substances (PFAS); Scoping Review.
Copyright © 2025 The Author(s). Published by Elsevier Ltd. All rights reserved.
Abstract
In this work, we are motivated by the problem of estimating racial disparities in health outcomes, specifically the average controlled difference (ACD) in telomere length between Black and White individuals, using data from the National Health and Nutrition Examination Survey (NHANES). To do so, we build a propensity for race to properly adjust for other social determinants while characterizing the controlled effect of race on telomere length. Propensity score methods are broadly employed with observational data as a tool to achieve covariate balance, but how to implement them in complex surveys is less studied-in particular, when the survey weights depend on the group variable under comparison (as the NHANES sampling scheme depends on self-reported race). We propose identification formulas to properly estimate the ACD in outcomes between Black and White individuals, with appropriate weighting for both covariate imbalance across the two racial groups and generalizability. Via extensive simulation, we show that our proposed methods outperform traditional analytic approaches in terms of bias, mean squared error, and coverage when estimating the ACD for our setting of interest. In our data, we find that evidence of racial differences in telomere length between Black and White individuals attenuates after accounting for confounding by socioeconomic factors and utilizing appropriate propensity score and survey weighting techniques. Software to implement these methods and code to reproduce our results can be found in the R package svycdiff, available through the Comprehensive R Archive Network (CRAN) at cran.r-project.org/web/packages/svycdiff/, or in a development version on GitHub at github.com/salernos/svycdiff.
Keywords: NHANES; complex surveys; controlled outcome differences; propensity scores; racial disparities.
© 2025 John Wiley & Sons Ltd.
Abstract
Background: Hepatocellular carcinoma (HCC) is among the most prevalent and deadly cancers worldwide. Chronic liver disease is the most established Risk factor for HCC, but Environmental Exposures are increasingly recognized as contributor. In this narrative review, we sought to analyze data linking three substances to HCC: polycyclic aromatic hydrocarbons (PAHs), Per- and Polyfluoroalkyl Substances (PFAS), and Cadmium. Methods: We performed a literature search of MEDLINE for this narrative review. Two reviewers screened titles and abstracts for relevance. We focused on articles published within the last three years. Results: PAH, PFAS, and Cadmium have been linked to chronic liver disease, liver injury, and to a lesser extent, HCC. Major limitations in existing data are small sample sizes, lack of longitudinal follow up (most studies are cross-sectional), and heterogeneity in the species assayed and the methods of assay. Conclusions: Further studies in large, prospective cohorts with longitudinal follow up are needed. Given existing evidence linking these substances to liver injury and HCC, a multi-faceted prevention and intervention strategy is needed, incorporating research, public education and engagement, legal frameworks, decontamination and medical interventions to mitigate deleterious effects of these substances.
Keywords: DNA damage; Oxidative stress; PFAS; Polycyclic aromatic hydrocarbons; Toxins.
© 2025. The Author(s), under exclusive licence to Springer Science+Business Media, LLC, part of Springer Nature.
Abstract
Background: The impact of short-chain, low molecular weight Polyfluoroalkyl substances (PFAS) and long-chain, high molecular weight PFAS on elevated alanine aminotransferase (ALT) remains unclear. Additionally, demographic and behavioral factors influencing PFAS levels in the U.S. population are not well understood. Objectives: To examine (1) associations between short- and long-chain PFAS mixtures and elevated ALT, and (2) participant characteristics linked to PFAS levels. Methods: This study included 378 adults (196 with detailed liver disease evaluations and 135 with cirrhosis), aged 40-75 years, from the Southern Liver Health Study (STRIVE), an ongoing prospective Cohort. Logistic regression assessed associations between serum PFAS and elevated ALT. Quantile g-computation evaluated PFAS mixture effects, while linear regression identified demographic and behavioral factors linked to PFAS levels. Results: A one-quartile increase in short-chain PFAS mixture levels was associated with higher odds of elevated ALT (adjusted odds ratio [aOR], 95 % confidence interval [CI]: 2.14, 1.31-3.50; P = 0.002) and higher log-transformed ALT levels (adjusted β, 95 % CI: 0.09, 0.01-0.17; P = 0.02). Long-chain PFAS mixtures showed no significant association. Individually, perfluoroheptanoic acid (a short-chain PFAS) (aOR, 95 % CI: 1.51, 1.001-2.27; P = 0.049) and perfluorohexanesulfonic acid (a long-chain PFAS) (aOR, 95 % CI: 1.53, 1.08-2.15; P = 0.015) were positively associated with elevated ALT. PFAS levels were lower in bottled water users but higher in current alcohol users. Males and older individuals exhibited higher long-chain PFAS levels, whereas non-water drinkers and current everyday smokers had lower levels of long-chain PFAS. Discussion: Higher short-chain PFAS mixture levels were linked to elevated ALT, with individual PFAS levels varying by sex and lifestyle factors. Limiting short-chain PFAS exposure may help prevent liver injury.
Keywords: Alanine transaminase; Fluorocarbons; Liver cirrhosis; Liver diseases; [MeSH].
Copyright © 2025. Published by Elsevier Inc.
Abstract
Objectives: To conduct a scoping review (ScR) of existing approaches for synthetic Electronic Health Records (EHR) data generation, to benchmark major methods, and to provide an open-source software and offer recommendations for practitioners. Materials and methods: We search three academic databases for our scoping review. Methods are benchmarked on open-source EHR datasets, Medical Information Mart for Intensive Care III and IV (MIMIC-III/IV). Seven existing methods covering major categories and two baseline methods are implemented and compared. Evaluation metrics concern data fidelity, downstream utility, privacy protection, and computational cost. Results: Forty-eight studies are identified and classified into five categories. Seven open-source methods covering all categories are selected, trained on MIMIC-III, and evaluated on MIMIC-III or MIMIC-IV for transportability considerations. Among them, Generative Adversarial Network (GAN)-based methods demonstrate competitive performance in fidelity and utility on MIMIC-III, rule-based methods excel in privacy protection. Similar findings are observed on MIMIC-IV, except that GAN-based methods further outperform the baseline methods in preserving fidelity. Discussion: Method choice is governed by the relative importance of the evaluation metrics in downstream use cases. We provide a decision tree to guide the choice among the benchmarked methods. An extensible Python package, “SynthEHRella”, is provided to facilitate streamlined evaluations. Conclusion: GAN-based methods excel when distributional shifts exist between the training and testing populations. Otherwise, CorGAN and MedGAN are most suitable for association modeling and predictive modeling, respectively. Future research should prioritize enhancing fidelity of the synthetic data while controlling privacy exposure, and comprehensive benchmarking of longitudinal or conditional generation methods.
Keywords: benchmarking; confidentiality; generative AI; scoping review; synthetic EHR.
© The Author(s) 2025. Published by Oxford University Press on behalf of the American Medical Informatics Association. All rights reserved. For commercial re-use, please contact [email protected] for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact [email protected].
Abstract
Background: Body dissatisfaction can drive individuals to use personal care products, exposing themselves to Benzophenone-3 (BP3). Yet, no study has examined the link between body dissatisfaction and elevated chemical exposures. Objectives: Our study examines how body dissatisfaction impacts the racial differences in BP3 exposures. Methods: Using NHANES 2003-2016 data for 3,072 women, we ascertained body dissatisfaction with a questionnaire on weight perception. We ran two generalized linear models with log10-transformed urinary concentrations of BP3 as the outcome variable and the following main predictors: one with race/ethnicity and another combining race/ethnicity and body dissatisfaction. We also conducted stratified analyses by race/ethnicity. We adjusted for poverty income ratio, BMI, urinary creatinine, and sunscreen usage. Results: BP3 levels in Mexican American, Other Hispanic, Other Race, non-Hispanic White, and non-Hispanic Asian women were on average 59 %, 56 %, 33 %, 16 %, and 9 % higher, respectively, compared to non-Hispanic Black women. Racial differences in BP3 levels are further widen with body dissatisfaction. For example, Other Hispanic women perceiving themselves as overweight had 69 % higher BP3 levels than non-Hispanic Black women (p-value = 0.01), while those perceiving themselves as at the right weight had 32 % higher levels (p-value = 0.31). Moreover, minority women perceiving themselves as overweight tended to have higher BP3 levels than those who do not. For example, BP3 levels in Other Hispanic women perceiving themselves as overweight are significantly higher compared to those who do not (73 %, p-value = 0.03). In contrast, such differences in the non-Hispanic White women are minimal (-0.5 %, p-value = 0.98). Discussion: Minority women with body dissatisfaction show elevated BP3 exposure independent of sunscreen usage, implying that their elevated exposures may stem from using other personal care and consumer products. Further research is needed to determine if increased exposure to other toxicants occur among minority women with body dissatisfaction.
Keywords: Body dissatisfaction; Body image; Chemical biomonitoring; Chemical exposures; Environmental justice; Racial disparities.
Copyright © 2025 The Author(s). Published by Elsevier Ltd. All rights reserved.
Abstract
Colorectal Cancer is highly preventable with timely screening, but screening modalities are widely underused, especially among those of low individual-level socioeconomic status (SES). In addition to individual-level SES, neighborhood-level SES may also play a role in colorectal Cancer screening completion through less geographic access to health care, transportation, and community knowledge of and support for screenings. We investigated the associations of neighborhood SES using a census tract-level measure of social and economic conditions with the uptake of colonoscopy and stool-based testing. We utilized data from the Southern Community Cohort Study, a large, prospective study of English-speaking adults ages 40 to 79 from the southeastern United States with 65% of participants identifying as non-Hispanic Black and 53% having annual household income <$15,000. Neighborhood SES was measured via a neighborhood deprivation index compiled from principal component analysis of 11 census-tract variables in the domains of education, employment, occupation, and poverty; screening was self-reported at the baseline interview (2002-2009) and follow-up interview (2008-2012). We found that participants residing in the lowest SES areas had lower odds of ever undergoing colonoscopy (ORQ5vsQ1 = 0.75; 95% confidence interval, 0.68-0.82) or stool-based colorectal Cancer testing (ORQ5vsQ1 = 0.71; 95% confidence interval, 0.63-0.80) while adjusting for individual-level SES factors. Associations were consistent between neighborhood SES and screening in subgroups defined by race, sex, household income, insurance, or education (P > 0.20 for all interaction tests). Our findings suggest that barriers to screening exist at the neighborhood level and that residents of lower SES neighborhoods may experience more barriers to screening using colonoscopy and stool-based modalities. Prevention Relevance: This study presents evidence that persons living in lower SES neighborhoods use colorectal Cancer screening modalities at lower rates. Screening is highly preventive of colorectal Cancer, but it has limited benefit if it cannot be utilized. Addressing neighborhood-level barriers to screening may improve socioeconomic disparities in colorectal Cancer.
©2025 American Association for Cancer Research.
Abstract
Background: Quantitative characterization of the health impacts associated with exposure to chemical mixtures has received considerable attention in current environmental and epidemiological studies. With many existing statistical methods and emerging approaches, it is important for practitioners to understand which method is best suited for their inferential goals. Objective: The goal of this paper is to provide empirical simulation-based evidence regarding performance of mixture methods to help guide researchers on selecting the best available methods to address three scientific questions in mixtures analysis: identifying important components of a mixture, identifying interactions among mixture components, and creating a summary score for risk stratification and prediction. Methods: We conducted a review and comparison of 11 analytical methods available for use in mixtures research through extensive simulation studies for continuous and binary outcomes. In addition, we carried out an illustrative data analysis using the PROTECT birth Cohort from Puerto Rico to examine the associations between exposure to chemical mixtures-metals, polycyclic aromatic hydrocarbons (PAHs), phthalates, and phenols-and birth outcomes. Results: Our simulation results suggest that the choice of methods depends on the goal of analysis and that there is no clear winner across the board. For selection of important toxicants in the mixtures and for identifying interactions, Elastic net (Enet) by Zou et al., Lasso for Hierarchical Interactions (HierNet) by Bien et al., and selection of nonlinear interactions by a forward stepwise algorithm (SNIF) by Narisetty et al. have the most stable performance across simulation settings. For overall summary or a cumulative measure, we find that using the Super Learner to combine multiple environmental risk scores can Lead to improved risk stratification and prediction properties. Conclusions: We develop an integrated R package “CompMix” that provides a platform for mixtures analysis where the practitioners can implement a pipeline that includes several approaches for mixtures analysis. Our study offers guidelines for selecting appropriate statistical methods for addressing specific scientific questions related to mixtures research. We identify critical gaps where new and better methods are needed. https://doi.org/10.1289/EHP15305.
Abstract
Propensity scores are commonly used to reduce the confounding bias in non-randomized observational studies for estimating the average treatment effect. An important assumption underlying this approach is that all confounders that are associated with both the treatment and the outcome of interest are measured and included in the propensity score model. In the absence of strong prior knowledge about potential confounders, researchers may agnostically want to adjust for a high-dimensional set of pre-treatment variables. As such, variable selection procedure is needed for propensity score estimation. In addition, studies show that including variables related to treatment only in the propensity score model may inflate the variance of the treatment effect estimators, while including variables that are predictive of only the outcome can improve efficiency. In this article, we propose to incorporate outcome-covariate relationship in the propensity score model by including the predicted binary outcome probability as a covariate. Our approach can be easily adapted to an ensemble of variable selection methods, including regularization methods and modern machine-learning tools based on classification and regression trees. We evaluate our method to estimate the treatment effects on a binary outcome, which is possibly censored, across multiple treatment groups. Simulation studies indicate that incorporating outcome probability for estimating the propensity scores can improve statistical efficiency and protect against model misspecification. The proposed methods are applied to a Cohort of advanced-stage prostate Cancer patients identified from a private insurance claims database for comparing the adverse effects of four commonly used drugs for treating castration-resistant prostate Cancer.
Keywords: Causal inference; claims data; confounder selection; high-dimensional; outcome-adaptive; propensity scores; tree-based methods.
Research Letter
Associations between adverse childhood experiences (ACEs) and liver disease in adulthood have been largely attributed to risk behaviors for viral and alcohol-related liver disease (ALD), as described in the landmark 1998 ACEs Study. Given the rise of metabolic dysfunction-associated steatohepatitis (MASH), we aim to update associations between ACEs and cirrhosis. This study included participants with and without cirrhosis aged 40-75 years living in North Carolina and Georgia enrolled in the Southern Liver Heath Study (STRIVE). Demographic data were collected via self-report and medical record review. History of 8 ACEs relating to abuse and household dysfunction (HD) were collected via validated survey. We tested the association between ACEs and cirrhosis using multivariable logistic regression models. See Supplementary Methods for additional details regarding study design. 461 participants were included (cirrhosis, n=187; non-cirrhosis, n=274; Table 1). Compared to non-cirrhosis participants, those with cirrhosis were more often male (46% vs. 28%), older (median 61.0 vs. 58.5 years), and non-Hispanic White (79% vs. 61%). Annual household income (HHI) was low in both groups with most households reporting less than $50,000. Participants with cirrhosis were significantly more likely to have diabetes (39.4% vs. 18.0%, p<0.001) and not currently use alcohol (12.9% vs. 52.3%, p<0.001). There were significant differences in the proportions of participants with cirrhosis enrolled at each recruiting site (p<0.001) by design. Among participants enrolled from Duke sites, the most common cirrhosis etiology was MASH (49.5%), followed by ALD (24.2%). Survey response rate was 49.5% in participants with cirrhosis and 62.5% for those without. A median of 1 ACE was reported in both cohorts (Supplementary Table 1). Cirrhosis participants more frequently reported ≥4 ACEs (20.3% vs. 15.3%), but in the adjusted model, wide confidence intervals made this estimate unstable [(adjusted OR (aOR) 1.67; 95% CI 0.73-3.82). 47.6% of participants with cirrhosis and 45.3% without cirrhosis reported experiencing childhood abuse (aOR 1.09; 95% CI 0.63-1.88), and the majority of participants with (62.0%) and without cirrhosis (56.2%) reported childhood HD. In the adjusted model, there was a significant difference such that participants with history of ≥1 ACE related to HD had greater odds of cirrhosis than those without ACEs related to HD (aOR 1.78; 95% CI 1.01-3.12). Additionally, we found a significant interaction between number of ACEs and HHI on the odds of cirrhosis, such that among participants with a HHI of less than $50,000 annually, those with ≥4 ACEs had 3 times the odds of cirrhosis versus those with 0-3 ACEs (p=0.041; Supplementary Table 2). Besides HHI, no other characteristics significantly interacted with the relationship between cirrhosis and total ACE score. There was no significant difference in total ACE score between the cirrhosis and non-cirrhosis cohorts (unadjusted OR 1.06; 95% CI 0.96-1.18), nor were there statistically significant differences in prevalence of any individual ACE between cohorts. Overall, our findings suggest that the association between exposure to ACEs and cirrhosis has likely changed in the decades since publication of the landmark ACEs Study, as we did not find a significant association between total ACE score and cirrhosis. However, we did find that HD was significantly associated with cirrhosis, suggesting that these exposures and their accompanying coping mechanisms may be more likely to contribute to cirrhosis than other ACEs. Prior studies have reported HD ACEs to be stronger predictors of both diabetes and dyslipidemia compared to other ACEs. Considering the high prevalence of MASH and its high comorbidity with diabetes and dyslipidemia, these studies support a potential shared exposure for MASH cirrhosis via these early household life stressors. Our results also highlighted an important interplay with HHI. Among participants who have low HHI, those with a high total ACE score have more than three times the odds of cirrhosis. This income bracket includes more than one-third of participants in both the cirrhosis and non-cirrhosis groups in our study. Concordantly, a previous study found that lower income was strongly correlated with higher mortality for Americans in the lowest 30% of income distribution. Low income has also been associated with increased prevalence of risk factors for cirrhosis, including obesity and alcohol misuse. Given that we found no association between ACEs and cirrhosis in higher income groups, there may be an income threshold that affords people sufficient access to healthcare and healthy lifestyles, below which the lack of health resources and other stressors associated with low HHI may compound ACEs. Future research is needed to understand the process by which ACEs promote disease. Emerging data suggest that epigenetic changes consistent with accelerated aging interact both with ACEs and later life disease. A recent study by Kim et. al. found that among middle-aged adults, having >=4 ACEs was associated with epigenetic age acceleration via alterations in DNA methylation, even after controlling for demographics, behavior and socioeconomic status. These findings when combined with ours suggest that early adversity may be associated with lasting changes in the biological aging process, and that this may increase risk for chronic diseases later in life. Limitations of the study include small sample size and small numbers of Hispanic participants. Recruitment methods also differed for the cirrhosis and non-cirrhosis cohorts. Participants with cirrhosis were recruited mainly from academic centers, whereas those without cirrhosis came primarily from community sites. This difference likely contributed to the differing participant characteristics between the cohorts. We attempted to control for these potential confounders by adjusting for them in our statistical modeling. Lastly, we did not have access to medical records for the non-cirrhosis Cohort, and it is possible that some may have undiagnosed liver disease, thus reducing power to detect an association. Considering that participants with cirrhosis were primarily enrolled from academic centers in urban areas in the Southeast, it may be worthwhile to determine whether our findings are generalizable to broader populations as well. Given the large global burden of liver disease, consideration of non-traditional risk factors such as ACEs will be essential in mitigating its future growth. Strategies are needed to prevent ACEs, as well as to lessen their impact on chronic disease later in life.
Abstract
Background: Clinical risk prediction models integrated into digitized health care informatics systems hold promise for personalized primary prevention and care, a core goal of precision health. Fairness metrics are important tools for evaluating potential disparities across sensitive features, such as sex and race or ethnicity, in the field of prediction modeling. However, fairness metric usage in clinical risk prediction models remains infrequent, sporadic, and rarely empirically evaluated. Objective: We seek to assess the uptake of fairness metrics in clinical risk prediction modeling through an empirical evaluation of popular prediction models for 2 diseases, 1 chronic and 1 infectious disease. Methods: We conducted a scoping literature review in November 2023 of recent high-impact publications on clinical risk prediction models for cardiovascular disease (CVD) and COVID-19 using Google Scholar. Results: Our review resulted in a shortlist of 23 CVD-focused articles and 22 COVID-19 pandemic-focused articles. No articles evaluated fairness metrics. Of the CVD-focused articles, 26% used a sex-stratified model, and of those with race or ethnicity data, 92% had study populations that were more than 50% from 1 race or ethnicity. Of the COVID-19 models, 9% used a sex-stratified model, and of those that included race or ethnicity data, 50% had study populations that were more than 50% from 1 race or ethnicity. No articles for either disease stratified their models by race or ethnicity. Conclusions: Our review shows that the use of fairness metrics for evaluating differences across sensitive features is rare, despite their ability to identify inequality and flag potential gaps in prevention and care. We also find that training data remain largely racially and ethnically homogeneous, demonstrating an urgent need for diversifying study cohorts and data collection. We propose an implementation framework to initiate change, calling for better connections between theory and practice when it comes to the adoption of fairness metrics for clinical risk prediction. We hypothesize that this integration will Lead to a more equitable prediction world.
Keywords: COVID-19; bias; cardiovascular disease; clinical risk prediction; equity; risk stratification; sensitive features.
©Lillian Rountree, Yi-Ting Lin, Chuyu Liu, Maxwell Salvatore, Andrew Admon, Brahmajee Nallamothu, Karandeep Singh, Anirban Basu, Fan Bu, Bhramar Mukherjee. Originally published in the Online Journal of Public Health Informatics (https://ojphi.jmir.org/), 19.03.2025.
Abstract
This report describes a comparison of two geocoding methods used by the Cohorts for Environmental Exposures and Cancer Risks (CEECR) Consortium: ArcGIS Geocoding by Esri and the SAS GEOCODE Procedure. The goal of this report is to determine the comparability of data sets that employ different approaches for linking survey data with spatial surrogates of exposure to environmental and socioeconomic factors. ArcGIS and SAS GEOCODE were selected as two platforms for this comparison because both programs are being used by one or more CEECR Cohort study teams and they can be used locally offline. The latter minimizes confidentiality issues related to online data linkages. Residential addresses from 3,238 Wisconsin residents in the Cancer & COVID Study and the Wisconsin in situ Cohort (WISC) were geocoded and linked to eight different publicly available datasets of environmental and socioeconomic factors at various geographic scales using both geocoding platforms. Since the two analytic platforms vary in geocoding approaches, the validity and accuracy of both platforms were compared to examine differences when assigning surrogate measurements of exposure based on spatial locations. ArcGIS offered a higher specificity for matched addresses with slightly more latitude/longitude point and street matches (97.7%) than SAS (95.9%), with the remainder matching at the zip code level. The two geocoding platforms showed high concordance in assignment at the county (99.6%), census tract (96.5%), and census block group (94.7%). As a result, the correlations based on census tracts and block groups were very strong for linked exposure measures of socioeconomic status, environmental justice, urban/rural residence, air pollution, proximity to industrial sites, and Cancer risk (all intraclass correlation coefficients ≥98%). Slightly lower concordance was observed for point source linkages (intraclass correlation coefficients 96-97%). Approximately ~4% of addresses were mis-matched largely in rural areas where census areas are larger and accurate geocoding base-layers are less widely available than in urban areas. For researchers that are already utilizing SAS, the GEOCODE procedure can be a logical choice as it is included in base SAS software and does not require an additional cost. However, SAS and ArcGIS provide similar options for the vast majority of study address locations.
Copyright © 2025 The Author(s).
Abstract
Factors driving accelerated biological age (BA), an important predictor of chronic diseases, remain poorly understood. This study focuses on the impact of diet and gut microbiome on accelerated BA. Accelerated Klemera-Doubal biological age (KDM-BA) was estimated as the difference between KDM-BA and chronological age. We assessed the cross-sectional association between accelerated KDM-BA and diet/gut microbiome in 117 adult participants from the 10,000 Families Study. 16S rRNA sequencing was used to estimate the abundances of gut bacterial genera. Multivariable linear mixed models evaluated the associations between accelerated KDM-BA and diet/gut microbiome after adjusting for family relatedness, diet, age, sex, smoking status, alcohol intake, and BMI. One standard deviation (SD) increase in processed meat was associated with a 1.91-year increase in accelerated KDM-BA (p = 0.04), while one SD increase in fiber intake was associated with a 0.70-year decrease in accelerated KDM-BA (p = 0.01). Accelerated KDM-BA was positively associated with Streptococcus and negatively associated with Subdoligranulum, unclassified Bacteroidetes, and Burkholderiales. Adjustment for gut microbiome did not change the association between dietary fiber and accelerated KDM-BA, but the association with processed meat intake became nonsignificant. These cross-sectional associations between higher meat intake, lower fiber intake, and accelerated BA need validation in longitudinal studies.
Keywords: accelerated aging; diet; gut microbiome.
Abstract
The human gut microbiome, the host, and the environment are inextricably linked across the life course with significant health impacts. Consisting of trillions of bacteria, fungi, viruses, and other micro-organisms, microbiota living within our gut are particularly dynamic and responsible for digestion and metabolism of diverse classes of ingested chemical pollutants. Exposure to chemical pollutants not only in early life but throughout growth and into adulthood can alter human hosts’ ability to absorb and metabolize xenobiotics, nutrients, and other components critical to health and longevity. Inflammation is a common mechanism underlying multiple environmentally related chronic conditions, including cardiovascular disease, multiple Cancer types, and mental health. While growing research supports complex interactions between pollutants and the gut microbiome, significant gaps exist. Few reviews provide descriptions of the complex mechanisms by which chemical pollutants interact with the host microbiome through either direct or indirect pathways to alter disease risk, with a particular focus on inflammatory pathways. This review focuses on examples of several classes of pollutants commonly ingested by humans, including (i) heavy metals, (ii) persistent organic pollutants (POPs), and (iii) nitrates. Digestive enzymes and gut microbes are the first line of absorption and metabolism of these chemicals, and gut microbes have been shown to alter compounds from a less to more toxic state influencing subsequent distribution and excretion. In addition, chemical pollutants may interact with or alter the selection of more harmful and less commensal microbiota, leading to gut dysbiosis, and changes in receptor-mediated signaling pathways that alter the integrity and function of the gut intestinal tract. Arsenic, Cadmium, and Lead (heavy metals), influence the microbiome directly by altering different classes of bacteria, and subsequently driving Inflammation through metabolite production and different signaling pathways (LPS/TLR4 or proteoglycan/TLR2 pathways). POPs can alter gut microbial composition either directly or indirectly depending on their ability to activate key signaling pathways within the intestine (e.g., PCB-126 and AHR). Nitrates and nitrites’ effect on the gut and host may depend on their ability to be transformed to secondary and tertiary metabolites by gut bacteria. Future research should continue to support foundational research both in vitro, in vivo, and longitudinal population-based research to better identify opportunities for prevention, gain additional mechanistic insights into the complex interactions between environmental pollutants and the microbiome and support additional translational science.
Keywords: Chemical pollutants; Dysbiosis; Gut microbiome; Gut pathology; Immune response; Inflammation; Xenobiotics.
Copyright © 2024 The Authors. Published by Elsevier Ltd. All rights reserved.
Abstract
Background: Per- and Polyfluoroalkyl Substances (PFAS) include thousands of manufactured compounds with growing public health concerns due to their potential for widespread human exposure and adverse health outcomes. While PFAS contamination remains a significant concern, especially from ingestion of contaminated food and water, determinants of the variability in PFAS exposure among regional and statewide populations in the United States remains unclear. Objectives: The objective of this study was to leverage The Survey of the Health of Wisconsin (SHOW), the only statewide representative Cohort in the US, to assess and characterize the variability of PFAS exposure in a general population. Methods: This study sample included a sub-sample of 605 adult participants from the 2014-2016 tri-annual statewide representative sample. Geometric means for PFOS, PFOA, PFNA, PFHxS, PFPeS, PFHpA, and a summed measure of 38 analyzed serum PFAS were presented by demographic, diet, behavioral, and residential characteristics. Multivariate linear regression was used to determine significant predictors of serum PFAS after adjustment. Results: Overall, higher serum concentrations of long-chain PFAS were observed compared with short-chain PFAS. Older adults, males, and non-Hispanic White individuals had higher serum PFAS compared to younger adults, females, and non-White individuals. Eating caught fish in the past year was associated with elevated levels of several PFAS. Discussion: This is among the first studies to characterize serum PFAS among a representative statewide sample in Wisconsin. Both short- and long-chain serum PFAS were detectable for six prominent PFAS. Age and consumption of great lakes fish were the most significant predictors of serum PFAS. State-level PFAS biomonitoring is important for identifying high risk populations and informing state public health standards and interventions, especially among those not living near known contamination sites.
Keywords: Determinants; Epidemiology; PFAS; Population-based.
Published by Elsevier Inc.
Abstract
Environmental chemical exposures influence immune system functions, and humans are exposed to a wide range of chemicals, termed the chemical “exposome”. A comprehensive, discovery analysis of the associations of multiple chemical families with immune biomarkers is needed. In this study, we tested the associations between environmental chemical concentrations and immune biomarkers. We analyzed the United States cross-sectional National Health and Nutrition Examination Survey (NHANES, 1999-2018). Chemical biomarker concentrations were measured in blood or urine (196 chemicals, 17 chemical families). Immune biomarkers included counts of lymphocytes, neutrophils, monocytes, basophils, eosinophils, red blood cells, white blood cells, and mean corpuscular volume. We conducted separate survey-weighted, multivariable linear regressions of each log2-transformed chemical and immune measure, adjusted for relevant covariates. We accounted for multiple comparisons using a false discovery rate (FDR). Among 45,528 adult participants, the mean age was 45.7 years, 51.4% were female, and 69.3% were Non-Hispanic White. 71 (36.2%) chemicals were associated with at least one of the eight immune biomarkers. The most chemical associations (FDR<0.05) were observed with mean corpuscular volume (36 chemicals) and red blood cell counts (35 chemicals). For example, a doubling in the concentration of cotinine was associated with 0.16 fL (95% CI: 0.15, 0.17; FDR<0.001) increased mean corpuscular volume, and a doubling in the concentration of blood Lead was associated with 61,736 increased red blood cells per μL (95% CI: 54,335, 69,138; FDR<0.001). A wide variety of chemicals, such as metals and smoking-related compounds, were highly associated with immune system biomarkers. This environmental chemical-wide association study identified chemicals from multiple families for further toxicological, immunologic, and epidemiological investigation.
Keywords: Environmental chemicals; Environmental epidemiology; Immune system biomarkers; Lead exposure; Metals; White blood cells.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Abstract
Environmental Exposures are recognized risk factors for Cancer etiology, but substantial gaps in knowledge remain, particularly regarding associations with emerging chemical exposures, underlying biological mechanisms of disease, and risk to racial/ethnic minorities and understudied populations. Exposures are complex due to the numerous mixtures in the environment, fluctuations over time, and different routes of exposure, all of which affect understanding of Cancer risk. Additional gaps include the impact of exposures on carcinogenesis during critical biologically relevant life stages. To address these gaps, in September 2021, the National Cancer Institute (NCI) in collaboration with the National Institute of Environmental Health Sciences (NIEHS) launched the Cohorts for Environmental Exposures and Cancer Risk (CEECR) program. CEECR supports five new large-scale population-based prospective Cohort studies for six years to address the impacts of a broad range of Environmental Exposures on intermediate Cancer biomarkers and Cancer risk, leveraging innovative approaches and technologies for Recruitment and exposure measures. These Cohort studies range from 10,000 to 100,000 participants and are being conducted in California, Minnesota, Michigan, and several southeastern states in the United States. The CEECR program has great potential to fill gaps in our understanding of environmental factors and Cancer etiology across multiple populations.
Abstract
Objectives: To develop recommendations regarding the use of weights to reduce selection bias for commonly performed analyses using electronic health record (EHR)-linked biobank data. Materials and methods: We mapped diagnosis (ICD code) data to standardized phecodes from 3 EHR-linked biobanks with varying Recruitment strategies: All of Us (AOU; n = 244 071), Michigan Genomics Initiative (MGI; n = 81 243), and UK Biobank (UKB; n = 401 167). Using 2019 National Health Interview Survey data, we constructed selection weights for AOU and MGI to represent the US adult population more. We used weights previously developed for UKB to represent the UKB-eligible population. We conducted 4 common analyses comparing unweighted and weighted results. Results: For AOU and MGI, estimated phecode prevalences decreased after weighting (weighted-unweighted median phecode prevalence ratio [MPR]: 0.82 and 0.61), while UKB estimates increased (MPR: 1.06). Weighting minimally impacted latent phenome dimensionality estimation. Comparing weighted versus unweighted phenome-wide association study for colorectal Cancer, the strongest associations remained unaltered, with considerable overlap in significant hits. Weighting affected the estimated log-odds ratio for sex and colorectal Cancer to align more closely with national registry-based estimates. Discussion: Weighting had a limited impact on dimensionality estimation and large-scale hypothesis testing but impacted prevalence and association estimation. When interested in estimating effect size, specific signals from untargeted association analyses should be followed up by weighted analysis. Conclusion: EHR-linked biobanks should report Recruitment and selection mechanisms and provide selection weights with defined target populations. Researchers should consider their intended estimands, specify source and target populations, and weight EHR-linked biobank analyses accordingly.
Keywords: ICD codes; biobank; electronic health records; phenome; selection bias.
© The Author(s) 2024. Published by Oxford University Press on behalf of the American Medical Informatics Association. All rights reserved. For permissions, please email: [email protected].
Abstract
Using administrative patient-care data such as Electronic Health Records (EHR) and medical/pharmaceutical claims for population-based scientific research has become increasingly common. With vast sample sizes leading to very small standard errors, researchers need to pay more attention to potential biases in the estimates of association parameters of interest, specifically to biases that do not diminish with increasing sample size. Of these multiple sources of biases, in this paper, we focus on understanding selection bias. We present an analytic framework using directed acyclic graphs for guiding applied researchers to dissect how different sources of selection bias may affect estimates of the association between a binary outcome and an exposure (continuous or categorical) of interest. We consider four easy-to-implement weighting approaches to reduce selection bias with accompanying variance formulae. We demonstrate through a simulation study when they can rescue us in practice with analysis of real-world data. We compare these methods using a data example where our goal is to estimate the well-known association of Cancer and biological sex, using EHR from a longitudinal biorepository at the University of Michigan Healthcare system. We provide annotated R codes to implement these weighted methods with associated inference.
Keywords: Michigan Genomics Initiative; calibration; directed acyclic graphs; inverse probability weighting; nonprobability sample; poststratification.
© The Royal Statistical Society 2024. All rights reserved. For commercial re-use, please contact [email protected] for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact [email protected].
Abstract
The etiology of lung Cancer in never-smokers remains elusive, despite 15% of lung Cancer cases in men and 53% in women worldwide being unrelated to smoking. Here, we aimed to enhance our understanding of lung Cancer pathogenesis among never-smokers using untargeted Metabolomics. This nested case-control study included 395 never-smoking women who developed lung Cancer and 395 matched never-smoking Cancer-free women from the prospective Shanghai Women’s Health Study with 15,353 metabolic features quantified in pre-diagnostic plasma using liquid chromatography high-resolution mass spectrometry. Recognizing that metabolites often correlate and seldom act independently in biological processes, we utilized a weighted correlation network analysis to agnostically construct 28 network modules of correlated metabolites. Using conditional logistic regression models, we assessed the associations for both metabolic network modules and individual metabolic features with lung Cancer, accounting for multiple testing using a false discovery rate (FDR) < 0.20. We identified a network module of 121 features inversely associated with all lung Cancer (p = .001, FDR = 0.028) and lung adenocarcinoma (p = .002, FDR = 0.056), where lyso-glycerophospholipids played a key role driving these associations. Another module of 440 features was inversely associated with lung adenocarcinoma (p = .014, FDR = 0.196). Individual metabolites within these network modules were enriched in biological pathways linked to oxidative stress, and energy metabolism. These pathways have been implicated in previous Metabolomics studies involving populations exposed to known lung Cancer risk factors such as traffic-related air pollution and polycyclic aromatic hydrocarbons. Our results suggest that untargeted plasma Metabolomics could provide novel insights into the etiology and risk factors of lung Cancer among never-smokers.
Keywords: lung Cancer; Metabolomics; network analysis; never‐smokers; oxidative stress.
© 2024 UICC. This article has been contributed to by U.S. Government employees and their work is in the public domain in the USA.
Abstract
The field of environmental epigenetics is uniquely suited to investigate biologic mechanisms that have the potential to link stressors to health disparities. However, it is common practice in basic epigenetic research to treat race as a covariable in large data analyses in a way that can perpetuate harmful biases without providing any biologic insight. In this article, we i) propose that epigenetic researchers open a dialogue about how and why race is employed in study designs and think critically about how this might perpetuate harmful biases; ii) call for interdisciplinary conversation and collaboration between epigeneticists and social scientists to promote the collection of more detailed social metrics, particularly institutional and structural metrics such as levels of discrimination that could improve our understanding of individual health outcomes; iii) encourage the development of standards and practices that promote full transparency about data collection methods, particularly with regard to race; and iv) encourage the field of epigenetics to continue to investigate how social structures contribute to biological health disparities, with a particular focus on the influence that structural racism may have in driving these health disparities.
Keywords: data transparency; epigenetic mechanisms; health inequities; race; structural determinants of health.
Copyright © 2024 King, Lalwani, Mercado, Dolan, Frierson, Meyer and Murphy.
Abstract
There has been raging discussion and debate around the quality of COVID death data in South Asia. According to WHO, of the 5.5 million reported COVID-19 deaths from 2020-2021, 0.57 million (10%) were contributed by five low and middle income countries (LMIC) countries in the Global South: India, Pakistan, Bangladesh, Sri Lanka and Nepal. However, a number of excess death estimates show that the actual death toll from COVID-19 is significantly higher than the reported number of deaths. For example, the IHME and WHO both project around 14.9 million total deaths, of which 4.5-5.5 million were attributed to these five countries in 2020-2021. We focus our gaze on the COVID-19 performance of these five countries where 23.5% of the world population lives in 2020 and 2021, via a counterfactual lens and ask, to what extent the mortality of one LMIC would have been affected if it adopted the pandemic policies of another, similar country? We use a Bayesian semi-mechanistic model developed by Mishra et al. (2021) to compare both the reported and estimated total death tolls by permuting the time-varying reproduction number (Rt) across these countries over a similar time period. Our analysis shows that, in the first half of 2021, mortality in India in terms of reported deaths could have been reduced to 96 and 102 deaths per million compared to actual 170 reported deaths per million had it adopted the policies of Nepal and Pakistan respectively. In terms of total deaths, India could have averted 481 and 466 deaths per million had it adopted the policies of Bangladesh and Pakistan. On the other hand, India had a lower number of reported COVID-19 deaths per million (48 deaths per million) and a lower estimated total deaths per million (80 deaths per million) in the second half of 2021, and LMICs other than Pakistan would have lower reported mortality had they followed India’s strategy. The gap between the reported and estimated total deaths highlights the varying level and extent of under-reporting of deaths across the subcontinent, and that model estimates are contingent on accuracy of the death data. Our analysis shows the importance of timely public health intervention and vaccines for lowering mortality and the need for better coverage infrastructure for the death registration system in LMICs.
Copyright: © 2023 Kundu et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Epigenetic researchers often evaluate DNA methylation as a potential mediator of the effect of social/Environmental Exposures on a health outcome. Modern statistical methods for jointly evaluating many mediators have not been widely adopted. We compare seven methods for high-dimensional mediation analysis with continuous outcomes through both diverse simulations and analysis of DNAm data from a large multi-ethnic Cohort in the United States, while providing an R package for their seamless implementation and adoption. Among the considered choices, the best-performing methods for detecting active mediators in simulations are the Bayesian sparse linear mixed model (BSLMM) and high-dimensional mediation analysis (HDMA); while the preferred methods for estimating the global mediation effect are high-dimensional linear mediation analysis (HILMA) and principal component mediation analysis (PCMA). We provide guidelines for epigenetic researchers on choosing the best method in practice and offer suggestions for future methodological development.
Copyright: © 2023 Clark-Boucher et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Importance: Natural language processing tools, such as ChatGPT (generative pretrained transformer, hereafter referred to as chatbot), have the potential to radically enhance the accessibility of medical information for health professionals and patients. Assessing the safety and efficacy of these tools in answering physician-generated questions is critical to determining their suitability in clinical settings, facilitating complex decision-making, and optimizing health care efficiency. Objective: To assess the accuracy and comprehensiveness of chatbot-generated responses to physician-developed medical queries, highlighting the reliability and limitations of artificial intelligence-generated medical information. Design, setting, and participants: Thirty-three physicians across 17 specialties generated 284 medical questions that they subjectively classified as easy, medium, or hard with either binary (yes or no) or descriptive answers. The physicians then graded the chatbot-generated answers to these questions for accuracy (6-point Likert scale with 1 being completely incorrect and 6 being completely correct) and completeness (3-point Likert scale, with 1 being incomplete and 3 being complete plus additional context). Scores were summarized with descriptive statistics and compared using the Mann-Whitney U test or the Kruskal-Wallis test. The study (including data analysis) was conducted from January to May 2023. Main outcomes and measures: Accuracy, completeness, and consistency over time and between 2 different versions (GPT-3.5 and GPT-4) of chatbot-generated medical responses. Results: Across all questions (n = 284) generated by 33 physicians (31 faculty members and 2 recent graduates from residency or fellowship programs) across 17 specialties, the median accuracy score was 5.5 (IQR, 4.0-6.0) (between almost completely and complete correct) with a mean (SD) score of 4.8 (1.6) (between mostly and almost completely correct). The median completeness score was 3.0 (IQR, 2.0-3.0) (complete and comprehensive) with a mean (SD) score of 2.5 (0.7). For questions rated easy, medium, and hard, the median accuracy scores were 6.0 (IQR, 5.0-6.0), 5.5 (IQR, 5.0-6.0), and 5.0 (IQR, 4.0-6.0), respectively (mean [SD] scores were 5.0 [1.5], 4.7 [1.7], and 4.6 [1.6], respectively; P = .05). Accuracy scores for binary and descriptive questions were similar (median score, 6.0 [IQR, 4.0-6.0] vs 5.0 [IQR, 3.4-6.0]; mean [SD] score, 4.9 [1.6] vs 4.7 [1.6]; P = .07). Of 36 questions with scores of 1.0 to 2.0, 34 were requeried or regraded 8 to 17 days later with substantial improvement (median score 2.0 [IQR, 1.0-3.0] vs 4.0 [IQR, 2.0-5.3]; P < .01). A subset of questions, regardless of initial scores (version 3.5), were regenerated and rescored using version 4 with improvement (mean accuracy [SD] score, 5.2 [1.5] vs 5.7 [0.8]; median score, 6.0 [IQR, 5.0-6.0] for original and 6.0 [IQR, 6.0-6.0] for rescored; P = .002). Conclusions and relevance: In this cross-sectional study, chatbot generated largely accurate information to diverse medical queries as judged by academic physician specialists with improvement over time, although it had important limitations. Further research and model development are needed to correct inaccuracies and for validation.
Abstract
Background: Some hormonally active cancers have low survival rates, but a large proportion of their incidence remains unexplained. Endocrine disrupting chemicals may affect hormone pathways in the pathology of these cancers. Objective: To evaluate cross-sectional associations between Per- and Polyfluoroalkyl Substances (PFAS), phenols, and parabens and self-reported previous Cancer diagnoses in the National Health and Nutrition Examination Survey (NHANES). Methods: We extracted concentrations of 7 PFAS and 12 phenols/parabens and self-reported diagnoses of melanoma and cancers of the thyroid, breast, ovary, uterus, and prostate in men and women (≥20 years). Associations between previous Cancer diagnoses and an interquartile range increase in exposure biomarkers were evaluated using logistic regression models adjusted for key covariates. We conceptualized race as social construct proxy of structural social factors and examined associations in non-Hispanic Black, Mexican American, and other Hispanic participants separately compared to White participants. Results: Previous melanoma in women was associated with higher PFDE (OR:2.07, 95% CI: 1.25, 3.43), PFNA (OR:1.72, 95% CI: 1.09, 2.73), PFUA (OR:1.76, 95% CI: 1.07, 2.89), BP3 (OR: 1.81, 95% CI: 1.10, 2.96), DCP25 (OR: 2.41, 95% CI: 1.22, 4.76), and DCP24 (OR: 1.85, 95% CI: 1.05, 3.26). Previous ovarian Cancer was associated with higher DCP25 (OR: 2.80, 95% CI: 1.08, 7.27), BPA (OR: 1.93, 95% CI: 1.11, 3.35) and BP3 (OR: 1.76, 95% CI: 1.00, 3.09). Previous uterine Cancer was associated with increased PFNA (OR: 1.55, 95% CI: 1.03, 2.34), while higher ethyl paraben was inversely associated (OR: 0.31, 95% CI: 0.12, 0.85). Various PFAS were associated with previous ovarian and uterine cancers in White women, while MPAH or BPF was associated with previous breast Cancer among non-White women. Impact statement: Biomarkers across all exposure categories (phenols, parabens, and per- and poly- fluoroalkyl substances) were cross-sectionally associated with increased odds of previous melanoma diagnoses in women, and increased odds of previous ovarian Cancer was associated with several phenols and parabens. Some associations differed by racial group, which is particularly impactful given the established racial disparities in distributions of exposure to these chemicals. This is the first epidemiological study to investigate exposure to phenols in relation to previous Cancer diagnoses, and the first NHANES study to explore racial/ethnic disparities in associations between environmental phenol, paraben, and PFAS exposures and historical Cancer diagnosis.
© 2023. The Author(s).
Abstract
There is a growing need for flexible general frameworks that integrate individual-level data with external summary information for improved statistical inference. External information relevant for a risk prediction model may come in multiple forms, through regression coefficient estimates or predicted values of the outcome variable. Different external models may use different sets of predictors and the algorithm they used to predict the outcome Y given these predictors may or may not be known. The underlying populations corresponding to each external model may be different from each other and from the internal study population. Motivated by a prostate Cancer risk prediction problem where novel biomarkers are measured only in the internal study, this paper proposes an imputation-based methodology, where the goal is to fit a target regression model with all available predictors in the internal study while utilizing summary information from external models that may have used only a subset of the predictors. The method allows for heterogeneity of covariate effects across the external populations. The proposed approach generates synthetic outcome data in each external population, uses stacked multiple imputation to create a long dataset with complete covariate information. The final analysis of the stacked imputed data is conducted by weighted regression. This flexible and unified approach can improve statistical efficiency of the estimated coefficients in the internal study, improve predictions by utilizing even partial information available from models that use a subset of the full set of covariates used in the internal study, and provide statistical inference for the external population with potentially different covariate effects from the internal population.
Keywords: data integration; prediction models; stacked multiple imputation; synthetic data.
© 2023 The Authors. Biometrics published by Wiley Periodicals LLC on behalf of International Biometric Society.
Du J, Zhou X, Clark-Boucher D, Hao W, Liu Y, Smith JA, Mukherjee B. Methods for large-scale single mediator hypothesis testing: Possible choices and comparisons. Genet Epidemiol. 2023 Mar;47(2):167-184. doi: 10.1002/gepi.22510. Epub 2022 Dec 8. PMID: 36465006.
Abstract
Mediation hypothesis testing for a large number of mediators is challenging due to the composite structure of the null hypothesis, 0 ( : effect of the exposure on the mediator after adjusting for confounders; : effect of the mediator on the outcome after adjusting for exposure and confounders). In this paper, we reviewed three classes of methods for large-scale one at a time mediation hypothesis testing. These methods are commonly used for continuous outcomes and continuous mediators assuming there is no exposure-mediator interaction so that the product has a causal interpretation as the indirect effect. The first class of methods ignores the impact of different structures under the composite null hypothesis, namely, (1) (2) ; and (3) . The second class of methods weights the reference distribution under each case of the null to form a mixture reference distribution. The third class constructs a composite test statistic using the three p values obtained under each case of the null so that the reference distribution of the composite statistic is approximately . In addition to these existing methods, we developed the Sobel-comp method belonging to the second class, which uses a corrected mixture reference distribution for Sobel’s test statistic. We performed extensive simulation studies to compare all six methods belonging to these three classes in terms of the false positive rates (FPRs) under the null hypothesis and the true positive rates under the alternative hypothesis. We found that the second class of methods which uses a mixture reference distribution could best maintain the FPRs at the nominal level under the null hypothesis and had the greatest true positive rates under the alternative hypothesis. We applied all methods to study the mediation mechanism of DNA methylation sites in the pathway from adult socioeconomic status to glycated hemoglobin level using data from the Multi-Ethnic Study of Atherosclerosis (MESA). We provide guidelines for choosing the optimal mediation hypothesis testing method in practice and develop an R package medScan available on the CRAN for implementing all the six methods.
Keywords: agnostic mediation analysis; composite null hypothesis; indirect effect; mediation effect; multiple hypothesis testing.
© 2022 The Authors. Genetic Epidemiology published by Wiley Periodicals LLC.
Abstract
The National Health and Nutrition Examination Survey (NHANES) provides data on the health and environmental exposure of the non-institutionalized US population. Such data have considerable potential to understand how the environment and behaviors impact human health. These data are also currently leveraged to answer public health questions such as prevalence of disease. However, these data need to first be processed before new insights can be derived through large-scale analyses. NHANES data are stored across hundreds of files with multiple inconsistencies. Correcting such inconsistencies takes systematic cross examination and considerable efforts but is required for accurately and reproducibly characterizing the associations between the exposome and diseases. Thus, we developed a set of curated and unified datasets and accompanied code by merging 614 separate files and harmonizing unrestricted data across NHANES III (1988-1994) and Continuous (1999-2018), totaling 134,310 participants and 4,740 variables. The variables convey 1) demographic information, 2) dietary consumption, 3) physical examination results, 4) occupation, 5) questionnaire items (e.g., physical activity, general health status, medical conditions), 6) medications, 7) mortality status linked from the National Death Index, 8) survey weights, 9) environmental exposure biomarker measurements, and 10) chemical comments that indicate which measurements are below or above the lower limit of detection. We also provide a data dictionary listing the variables and their descriptions to help researchers browse the data. We also provide R markdown files to show example codes on calculating summary statistics and running regression models to help accelerate high-throughput analysis and secular trends of the exposome.
Clark-Boucher D, Boss J, Salvatore M, Smith JA, Fritsche LG, Mukherjee B. Assessing the added value of linking electronic health records to improve the prediction of self-reported COVID-19 testing and diagnosis. PLoS One. 2022 Jul 25;17(7):e0269017. doi: 10.1371/journal.pone.0269017. PMID: 35877617; PMCID: PMC9312965.
Abstract
Since the beginning of the Coronavirus Disease 2019 (COVID-19) pandemic, a focus of research has been to identify risk factors associated with COVID-19-related outcomes, such as testing and diagnosis, and use them to build prediction models. Existing studies have used data from digital surveys or electronic health records (EHRs), but very few have linked the two sources to build joint predictive models. In this study, we used survey data on 7,054 patients from the Michigan Genomics Initiative biorepository to evaluate how well self-reported data could be integrated with electronic records for the purpose of modeling COVID-19-related outcomes. We observed that among survey respondents, self-reported COVID-19 diagnosis captured a larger number of cases than the corresponding EHRs, suggesting that self-reported outcomes may be better than EHRs for distinguishing COVID-19 cases from controls. In the modeling context, we compared the utility of survey- and EHR-derived predictor variables in models of survey-reported COVID-19 testing and diagnosis. We found that survey-derived predictors produced uniformly stronger models than EHR-derived predictors-likely due to their specificity, temporal proximity, and breadth-and that combining predictors from both sources offered no consistent improvement compared to using survey-based predictors alone. Our results suggest that, even though general EHRs are useful in predictive models of COVID-19 outcomes, they may not be essential in those models when rich survey data are already available. The two data sources together may offer better prediction for COVID severity, but we did not have enough severe cases in the survey respondents to assess that hypothesis in in our study.
Nguyen VK, Colacino J, Patel CJ, Sartor M, Jolliet O. Identification of occupations susceptible to high exposure and risk associated with multiple toxicants in an observational study: National Health and Nutrition Examination Survey 1999-2014. Exposome. 2022 Jun 25;2(1):osac004. doi: 10.1093/exposome/osac004. PMID: 35832257; PMCID: PMC9266352.
Abstract
Occupational exposures to toxicants are estimated to cause over 370 000 premature deaths annually. The risks due to multiple workplace chemical exposures and those occupations most susceptible to the resulting health effects remain poorly characterized. The aim of this study is to identify occupations with elevated toxicant biomarker concentrations and increased health risk associated with toxicant exposures in a diverse working US population. For this observational study of 51 008 participants, we used data from the 1999-2014 National Health and Nutrition Examination Survey. We characterized differences in chemical exposures by occupational group for 131 chemicals by applying a series of generalized linear models with the outcome as biomarker concentrations and the main predictor as the occupational groups, adjusting for age, sex, race/ethnicity, poverty income ratio, study period, and biomarker of tobacco use. For each occupational group, we calculated percentages of participants with chemical biomarker levels exceeding acceptable health-based guidelines. Blue-collar workers from “Construction,” “Professional, Scientific, Technical Services,” “Real Estate, Rental, Leasing,” “Manufacturing,” and “Wholesale Trade” have higher biomarker levels of toxicants such as several heavy metals, Acrylamide, glycideamide, and several volatile organic compounds (VOCs) compared with their white-collar counterparts. Moreover, blue-collar workers from these industries have toxicant concentrations exceeding acceptable levels: Arsenic (16%-58%), Lead (1%-3%), Cadmium (1%-11%), glycideamide (3%-6%), and VOCs (1%-33%). Blue-collar workers have higher toxicant levels relative to their white-collar counterparts, often exceeding acceptable levels associated with noncancer effects. Our findings identify multiple occupations to prioritize for targeted interventions and health policies to monitor and reduce toxicant exposures.
Keywords: biomonitoring equivalents; environmental chemicals; occupational epidemiology; occupational exposures; risk assessment; unsupervised learning.
© The Author(s) 2022. Published by Oxford University Press.
Fuentes ZC, Schwartz YL, Robuck AR, Walker DI. Operationalizing the Exposome Using Passive Silicone Samplers. Curr Pollut Rep. 2022;8(1):1-29. doi: 10.1007/s40726-021-00211-6. Epub 2022 Jan 4. PMID: 35004129; PMCID: PMC8724229.
Abstract
The exposome, which is defined as the cumulative effect of Environmental Exposures and corresponding biological responses, aims to provide a comprehensive measure for evaluating non-genetic causes of disease. Operationalization of the exposome for environmental health and precision medicine has been limited by the lack of a universal approach for characterizing complex exposures, particularly as they vary temporally and geographically. To overcome these challenges, passive sampling devices (PSDs) provide a key measurement strategy for deep exposome phenotyping, which aims to provide comprehensive chemical assessment using untargeted high-resolution mass spectrometry for exposome-wide association studies. To highlight the advantages of silicone PSDs, we review their use in population studies and evaluate the broad range of applications and chemical classes characterized using these samplers. We assess key aspects of incorporating PSDs within observational studies, including the need to preclean samplers prior to use to remove impurities that interfere with compound detection, analytical considerations, and cost. We close with strategies on how to incorporate measures of the external exposome using PSDs, and their advantages for reducing variability in exposure measures and providing a more thorough accounting of the exposome. Continued development and application of silicone PSDs will facilitate greater understanding of how Environmental Exposures drive disease risk, while providing a feasible strategy for incorporating untargeted, high-resolution characterization of the external exposome in human studies.
Keywords: Exposome; Exposure assessment; High-resolution mass spectrometry; Precision medicine; Silicone wristband samplers.
© The Author(s) 2021.
Datta J, Mukherjee B. Discussion on “Regression Models for Understanding COVID-19 Epidemic Dynamics with Incomplete Data”. J Am Stat Assoc. 2021;116(536):1583-1586. doi: 10.1080/01621459.2021.1982721. Epub 2021 Dec 16. PMID: 39439741; PMCID: PMC11495649.
Abstract
No Abstract Available
Keywords: COVID-19; Effective reproduction number; Retrospective analysis; Seroprevalence; unascertained cases.
© The Author(s) 2021.